Structural Variation in Genomes
Tobias Marschall
Structural Variation in Genomes
Each of our body cells harbors a copy of our genome, the set of all genetic material. The genome is organized in chromosomes: we inherit 23 chromosomes from our mother and father, respectively. A chromosome consists of a long DNA molecule, stabilized and spatially structured by special proteins. The DNA encodes genetic information as a sequence of its constituting bases adenine (A), cytosine (C), guanine (G), and thymine (T). In this sense, a chromosome can be viewed as a sequence of the letters A, C, G, and T. Likewise, we can represent a genome as a set of such sequences (one for each chromosome). The research fi eld of genomics asks the question, among others, of how our genome infl uences our traits. Many different traits can be of interest, such as body height, the risk of contracting a certain disease, or the tolerance to a drug, for instance. The genomes of two persons can differ in a multitude of ways, as illustrated in fi gure 1. Traditionally, many genetic studies focused on point mutations (SNPs). That is, one restricts oneself to differences in single letters. In recent years, however, increasing attention has also been given to larger differences, the so-called structural variants, such as deletions or duplications of whole DNA segments. Such variants occur frequently in humans and, hence, studying them is indispensable for capturing the full spectrum of human genetic diversity. This is enabled by new sequencing technologies in combination with novel algorithmic methods, which we develop actively. High-throughput methods for genome sequencing allow for reading millions of short DNA fragments in parallel. That means, these devices do not return the sequence of a whole chromosome; instead, they output the sequences of short fragments, called reads. The analysis of the resulting data is thus comparable to solving a jigsaw puzzle: from the reads, we want to reconstruct the sequenced genome. Here, we particularly focus on detecting structural differences.
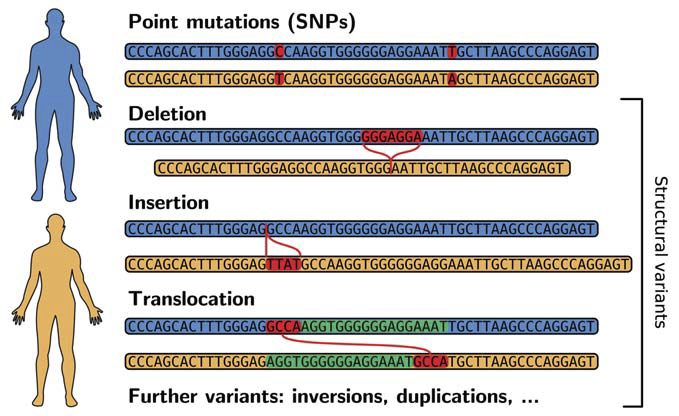
Differences between the genomic sequences of two persons
In a large study of 250 families, our computational methods have contributed to characterizing structural genetic variants in detail. In this project, the complete genomes of both parents and one child were sequenced for each family. The large number of studied families as well as the high sequencing depth render this effort a leading project worldwide. By reading the genomes of parents and children, we can determine which genetic variants found in the children were not inherited from the parents, but are the result of new mutations. The rate at which these so-called de novo mutations happen is of great interest as a parameter in models of human evolution.
Beyond fundamental research, studying structural genetic variants is potentially important for personalized medicine. So far, association studies that aim to establish connections between the genome of a person and clinically relevant traits were restricted to point mutations. Our recent research provides evidence that taking structural variants into account in such studies leads to new, medically relevant insights.
Tobias Marschall

DEPT. 3 Computational Biology and Applied Algorithmics
Phone +49 681 302 70880
Emailt.marschall mpi-inf.mpg.dehttp://mpi-inf.mpg.de
